Running individual simulation and retrieving the results
Once the simulation is loaded (see Loading a
simulation and accessing entities), it can be run using
runSimulations() to produce an object of the class
SimulationResults. Keep in mind that
runSimulations() produces a list of
SimulationResults objects.
library(ospsuite)
#> The option 'ospsuite.plots.watermarkEnabled' is not set.
#> To enable watermarks, add the following to your .Rprofile:
#> options(ospsuite.plots.watermarkEnabled = TRUE)
#> To disable watermarks, add:
#> options(ospsuite.plots.watermarkEnabled = FALSE)
#> You can edit your .Rprofile with usethis::edit_r_profile()
# Load the simulation
simFilePath <- system.file("extdata", "Aciclovir.pkml", package = "ospsuite")
sim <- loadSimulation(simFilePath)
simulationResults <- runSimulations(simulations = sim)
# Extract `SimulationResults` by simulation id
simulationResults <- simulationResults[[sim$id]]
print(simulationResults)
#> <SimulationResults>
#> • Number of individuals: 1
#> For paths:
#> • Organism|PeripheralVenousBlood|Aciclovir|Plasma (Peripheral Venous Blood)
#> • Organism|VenousBlood|Plasma|Aciclovir|Plasma UnboundThe advantage of storing the results in a object is the option to keep different results of the same simulation produced with different settings (e.g., model parameters).
Simulated time-value pairs for a specific output from the
SimulationResults-object can be accessed with the method
getOutputValues. The user can provide either the path(s) of
the output (which can be a molecule, a parameter, or an observer), or
the object(s) of the type Molecule, Parameter,
or Quantity (for observers) with the argument
quantitiesOrPaths. If no output is specified, all outputs
available in the simulation results are returned.
The paths of all available outputs can be accessed via
simulationResults$allQuantityPaths
#> [1] "Organism|PeripheralVenousBlood|Aciclovir|Plasma (Peripheral Venous Blood)"
#> [2] "Organism|VenousBlood|Plasma|Aciclovir|Plasma Unbound"getOutputValues returns a list with two entries:
data and metadata:
-
datais a dataframe with two predefined columns (IndividualId and Time) as well as one column for each requested output-
IndividualId(not relevant for an individual simulation) -
Timea vector with simulated time values (in minutes, equal for all outputs) - a vector with simulated entries for each output requested.
-
metaDatais a list containing one entry for each requested output. Each entry contains information pertinent to the output such as its dimension or the unit.
# Get simulated results by path
resultsPath <- simulationResults$allQuantityPaths[[1]]
print(resultsPath)
#> [1] "Organism|PeripheralVenousBlood|Aciclovir|Plasma (Peripheral Venous Blood)"
resultsData <- getOutputValues(
simulationResults,
quantitiesOrPaths = resultsPath
)
resultsTime <- resultsData$data$Time
resultsValues <- resultsData$data$`Organism|PeripheralVenousBlood|Aciclovir|Plasma (Peripheral Venous Blood)`
plot(resultsTime, resultsValues, type = "l")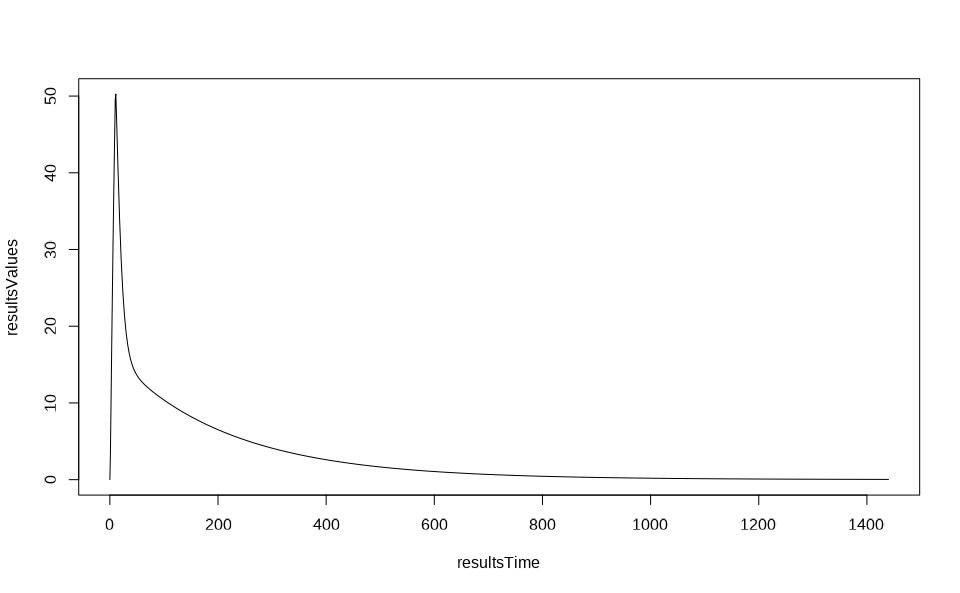
The results can be stored in and imported from a *.csv file with the
methods exportResultsToCSV and
importResultsFromCSV.
# Load and run the simulation
simFilePath <- system.file("extdata", "Aciclovir.pkml", package = "ospsuite")
sim <- loadSimulation(simFilePath)
# `runSimulations` returns a list of `SimulationResults`. For single simulation,
# we directly extract the first element, as only one object is created.
simulationResults <- runSimulations(simulations = sim)[[1]]
# Export to csv
csvResultsPath <- system.file("extdata", "SimResults.csv", package = "ospsuite")
exportResultsToCSV(results = simulationResults, filePath = csvResultsPath)
# Load from csv
resultsLoaded <- importResultsFromCSV(
filePaths = csvResultsPath,
simulation = sim
)
print(resultsLoaded)
#> <SimulationResults>
#> • Number of individuals: 1
#> For paths:
#> • Organism|PeripheralVenousBlood|Aciclovir|Plasma (Peripheral Venous Blood)
#> • Organism|VenousBlood|Plasma|Aciclovir|Plasma UnboundRunning multiple individual simulations and retrieving the results
In some cases, the user might want to run multiple simulations in
parallel. This can be achieved easily by simply passing a list of
simulations to the runSimulations() function. However, only
individual simulations (i.e., the population argument must
remain empty) are supported.
By default, the simulations will be executed in parallel by using up to all cores available on the machine minus 1. So if there are 8 cores and the user simulates 4 simulations, 4 cores will be used. On the other hand, if the user simulates 10 simulations with only 8 cores available, 7 will be used and then the first 3 available.
# Load and run multiple simulations concurrently.
simFilePath <- system.file("extdata", "Aciclovir.pkml", package = "ospsuite")
# We load 3 times the same simulation for convenience. But in real life scenarios,
# they should be different simulations
sim1 <- loadSimulation(simFilePath)
sim2 <- loadSimulation(simFilePath)
sim3 <- loadSimulation(simFilePath)
simulationResults <- runSimulations(simulations = list(sim1, sim2, sim3))The results in this case will be a named list of
SimulationResults-object , i.e. one for each simulation.
The names of the entries of the list are the ids of the simulation to
which the results correspond. This way, it is easy to retrieve the
correct results for the specific simulation
Adding new outputs
By default, only outputs that were selected in PK-Sim or MoBi prior
to the export of the simulation to pkml are generated. The
user can add new outputs to the simulation with the method
addOutputs. The outputs can be provided either as objects
of the type(s) Molecule, Parameter, or
Quantity, or as path strings. The output list is a property
of the simulation. After adding or removing outputs, the
corresponding simulation needs to be re-run in order to generate updated
results.
# Clear the list of generated outputs
clearOutputs(sim)
# Add new outputs as objects
molecule <- getMolecule("Organism|Kidney|Intracellular|Aciclovir", sim)
observer <- getQuantity("Organism|Lumen|Aciclovir|Fraction dissolved", sim)
addOutputs(c(molecule, observer), simulation = sim)
# Add new outputs as path strings
addOutputs(
c(
"Organism|Lumen|Stomach|Aciclovir",
"Organism|PeripheralVenousBlood|Aciclovir|Whole Blood (Peripheral Venous Blood)"
),
simulation = sim
)
# Run simulation
simulationResults <- runSimulations(simulations = sim)
# Retrieve all generated outputs (e.g. omitting the quantitiesOrPaths property
# will return all available values)
resultsData <- getOutputValues(simulationResults)
# Note that "Organism|PeripheralVenousBlood|Aciclovir|Plasma (Peripheral Venous Blood)"
# is not in the list of generated results any more
names(resultsData$data)
#> NULLclearOutputs() and addOutputs() can be
combined with the function setOutputs().
Changing simulation intervals
The simulation interval (i.e., the simulation times at which results
are stored) are stored as the property $outputSchema of a
Simulation object.
print(sim$outputSchema)
#> <OutputSchema>
#>
#> ── Output intervals ──
#>
#> <Interval>
#> • Name: Simulation Interval 1
#> • Start time: 0.00e+00 [min]
#> • End time: 15.00 [min]
#> • Resolution: 1.00 [pts/min]
#> <Interval>
#> • Name: Simulation Interval 2
#> • Start time: 15.00 [min]
#> • End time: 1440.00 [min]
#> • Resolution: 0.33 [pts/min]
#> <Interval>
#> • Name: Simulation Interval 3
#> • Start time: 120.00 [min]
#> • End time: 1440.00 [min]
#> • Resolution: 0.07 [pts/min]To change the simulation interval, the user can use one of the
functions clearOutputIntervals(),
addOutputInterval(), and
setOutputInterval().
# Remove all output intervals - simulation not possible!
clearOutputIntervals(simulation = sim)
runSimulations(simulations = sim)
#> Error in `do.call()`:
#> ! Type: OSPSuite.Utility.Exceptions.OSPSuiteException
#> Message: Time points output schema is empty
#> Method: Void EvaluateCppCallResult(Boolean, System.String)
#> Stack trace:
#> at OSPSuite.SimModel.PInvokeHelper.EvaluateCppCallResult(Boolean success, String errorMessage)
#> at OSPSuite.SimModel.Simulation.evaluateCppCallResult(Boolean success, String errorMessage)
#> at OSPSuite.SimModel.Simulation.RunSimulation()
#> at OSPSuite.Core.Domain.Services.SimModelManager.simulate()
#> at System.Threading.ExecutionContext.RunFromThreadPoolDispatchLoop(Thread threadPoolThread, ExecutionContext executionContext, ContextCallback callback, Object state)
#> --- End of stack trace from previous location ---
#> at System.Threading.ExecutionContext.RunFromThreadPoolDispatchLoop(Thread threadPoolThread, ExecutionContext executionContext, ContextCallback callback, Object state)
#> at System.Threading.Tasks.Task.ExecuteWithThreadLocal(Task& currentTaskSlot, Thread threadPoolThread)
#> --- End of stack trace fr
# Add an interval
addOutputInterval(
simulation = sim,
startTime = 0,
endTime = 20,
resolution = 60,
intervalName = "highRes"
)
print(sim$outputSchema)
#> <OutputSchema>
#>
#> ── Output intervals ──
#>
#> <Interval>
#> • Name: highRes
#> • Start time: 0.00e+00 [min]
#> • End time: 20.00 [min]
#> • Resolution: 60.00 [pts/min]
# Add a second interval
addOutputInterval(
simulation = sim,
startTime = 30,
endTime = 2000,
resolution = 4,
intervalName = "lowRes"
)
print(sim$outputSchema)
#> <OutputSchema>
#>
#> ── Output intervals ──
#>
#> <Interval>
#> • Name: highRes
#> • Start time: 0.00e+00 [min]
#> • End time: 20.00 [min]
#> • Resolution: 60.00 [pts/min]
#> <Interval>
#> • Name: lowRes
#> • Start time: 30.00 [min]
#> • End time: 2000.00 [min]
#> • Resolution: 4.00 [pts/min]
# Replace the existing interval(s) with a new one
setOutputInterval(
simulation = sim,
startTime = 0,
endTime = 2000,
resolution = 4
)
print(sim$outputSchema)
#> <OutputSchema>
#>
#> ── Output intervals ──
#>
#> <Interval>
#> • Name: Output interval
#> • Start time: 0.00e+00 [min]
#> • End time: 2000.00 [min]
#> • Resolution: 4.00 [pts/min]Adjusting solver settings
The solver settings control the numerical integration algorithm used
to solve the differential equations in the simulation. These settings
can be accessed and modified through the $solver property
of a Simulation object.
# View current solver settings
print(sim$solver)
#> <SolverSettings>
#> • useJacobian: TRUE
#> • h0: 1e-10
#> • hMin: 0
#> • hMax: 60
#> • mxStep: 100000
#> • relTol: 1e-05
#> • absTol: 1e-10
#> • checkForNegativeValues: TRUEIn some cases, a simulation may fail to run successfully due to numerical instability or convergence issues. One common approach to resolve such issues is to reduce the solver tolerances, which makes the solver use smaller step sizes and more precise calculations. The two main tolerance parameters are:
-
absTol: Absolute tolerance - controls the accuracy of the solution in absolute terms -
relTol: Relative tolerance - controls the accuracy of the solution relative to the magnitude of the values
# If a simulation fails to run, try reducing the tolerances
# Reduce absolute tolerance by a factor of 10
sim$solver$absTol <- sim$solver$absTol * 1e-1
# Optionally, the relative tolerance can also be reduced
# sim$solver$relTol <- sim$solver$relTol * 1e-1
# After changing solver settings, re-run the simulation
simulationResults <- runSimulations(simulations = sim)[[1]]Other solver settings that can be adjusted include:
-
h0: Initial time step size -
hMin: Minimum step size allowed -
hMax: Maximum step size allowed -
mxStep: Maximum number of (internal) steps to be taken by the solver in its attempt to reach the next output time -
useJacobian: Indicates whether to use the analytically calculated Jacobian matrix during calculations. When disabled, an approximation by difference quotients is used, which is much slower in the majority of cases
For more details on solver properties and their effects, see the MoBi documentation on ODE Solver Properties.